Parallel coordinate plot with information about missing/imputed values
Source:R/parcoordMiss.R
parcoordMiss.RdParallel coordinate plot with adjustments for missing/imputed values. Missing values in the plotted variables may be represented by a point above the corresponding coordinate axis to prevent disconnected lines. In addition, observations with missing/imputed values in selected variables may be highlighted.
parcoordMiss(
x,
delimiter = NULL,
highlight = NULL,
selection = c("any", "all"),
plotvars = NULL,
plotNA = TRUE,
col = c("skyblue", "red", "skyblue4", "red4", "orange", "orange4"),
alpha = NULL,
lty = par("lty"),
xlim = NULL,
ylim = NULL,
main = NULL,
sub = NULL,
xlab = NULL,
ylab = NULL,
labels = TRUE,
xpd = NULL,
interactive = TRUE,
...
)Arguments
- x
a matrix or
data.frame.- delimiter
a character-vector to distinguish between variables and imputation-indices for imputed variables (therefore,
xneeds to havecolnames()). If given, it is used to determine the corresponding imputation-index for any imputed variable (a logical-vector indicating which values of the variable have been imputed). If such imputation-indices are found, they are used for highlighting and the colors are adjusted according to the given colors for imputed variables (seecol).- highlight
a vector giving the variables to be used for highlighting. If
NULL(the default), all variables are used for highlighting.- selection
the selection method for highlighting missing/imputed values in multiple highlight variables. Possible values are
"any"(highlighting of missing/imputed values in any of the highlight variables) and"all"(highlighting of missing/imputed values in all of the highlight variables).- plotvars
a vector giving the variables to be plotted. If
NULL(the default), all variables are plotted.- plotNA
a logical indicating whether missing values in the plot variables should be represented by a point above the corresponding coordinate axis to prevent disconnected lines.
- col
if
plotNAisTRUE, a vector of length six giving the colors to be used for observations with different combinations of observed and missing/imputed values in the plot variables and highlight variables (vectors of length one or two are recycled). Otherwise, a vector of length two giving the colors for non-highlighted and highlighted observations (if a single color is supplied, it is used for both).- alpha
a numeric value between 0 and 1 giving the level of transparency of the colors, or
NULL. This can be used to prevent overplotting.- lty
if
plotNAisTRUE, a vector of length four giving the line types to be used for observations with different combinations of observed and missing/imputed values in the plot variables and highlight variables (vectors of length one or two are recycled). Otherwise, a vector of length two giving the line types for non-highlighted and highlighted observations (if a single line type is supplied, it is used for both).- xlim, ylim
axis limits.
- main, sub
main and sub title.
- xlab, ylab
axis labels.
- labels
either a logical indicating whether labels should be plotted below each coordinate axis, or a character vector giving the labels.
- xpd
a logical indicating whether the lines should be allowed to go outside the plot region. If
NULL, it defaults toTRUEunless axis limits are specified.- interactive
a logical indicating whether interactive features should be enabled (see ‘Details’).
- ...
for
parcoordMiss, further graphical parameters to be passed down (seegraphics::par()). ForTKRparcoordMiss, further arguments to be passed toparcoordMiss.
Details
In parallel coordinate plots, the variables are represented by parallel
axes. Each observation of the scaled data is shown as a line. Observations
with missing/imputed values in selected variables may thereby be
highlighted. However, plotting variables with missing values results in
disconnected lines, making it impossible to trace the respective
observations across the graph. As a remedy, missing values may be
represented by a point above the corresponding coordinate axis, which is
separated from the main plot by a small gap and a horizontal line, as
determined by plotNA. Connected lines can then be drawn for all
observations. Nevertheless, a caveat of this display is that it may draw
attention away from the main relationships between the variables.
If interactive is TRUE, it is possible switch between this
display and the standard display without the separate level for missing
values by clicking in the top margin of the plot. In addition, the variables
to be used for highlighting can be selected interactively. Observations
with missing/imputed values in any or in all of the selected variables are
highlighted (as determined by selection). A variable can be added to
the selection by clicking on a coordinate axis. If a variable is already
selected, clicking on its coordinate axis removes it from the selection.
Clicking anywhere outside the plot region (except the top margin, if
missing/imputed values exist) quits the interactive session.
Note
Some of the argument names and positions have changed with versions
1.3 and 1.4 due to extended functionality and for more consistency with
other plot functions in VIM. For back compatibility, the arguments
colcomb and xaxlabels can still be supplied to ...{}
and are handled correctly. Nevertheless, they are deprecated and no longer
documented. Use highlight and labels instead.
References
Wegman, E. J. (1990) Hyperdimensional data analysis using parallel coordinates. Journal of the American Statistical Association 85 (411), 664–675.
M. Templ, A. Alfons, P. Filzmoser (2012) Exploring incomplete data using visualization tools. Journal of Advances in Data Analysis and Classification, Online first. DOI: 10.1007/s11634-011-0102-y.
See also
Other plotting functions:
aggr(),
barMiss(),
histMiss(),
marginmatrix(),
marginplot(),
matrixplot(),
mosaicMiss(),
pairsVIM(),
pbox(),
scattJitt(),
scattMiss(),
scattmatrixMiss(),
spineMiss()
Examples
data(chorizonDL, package = "VIM")
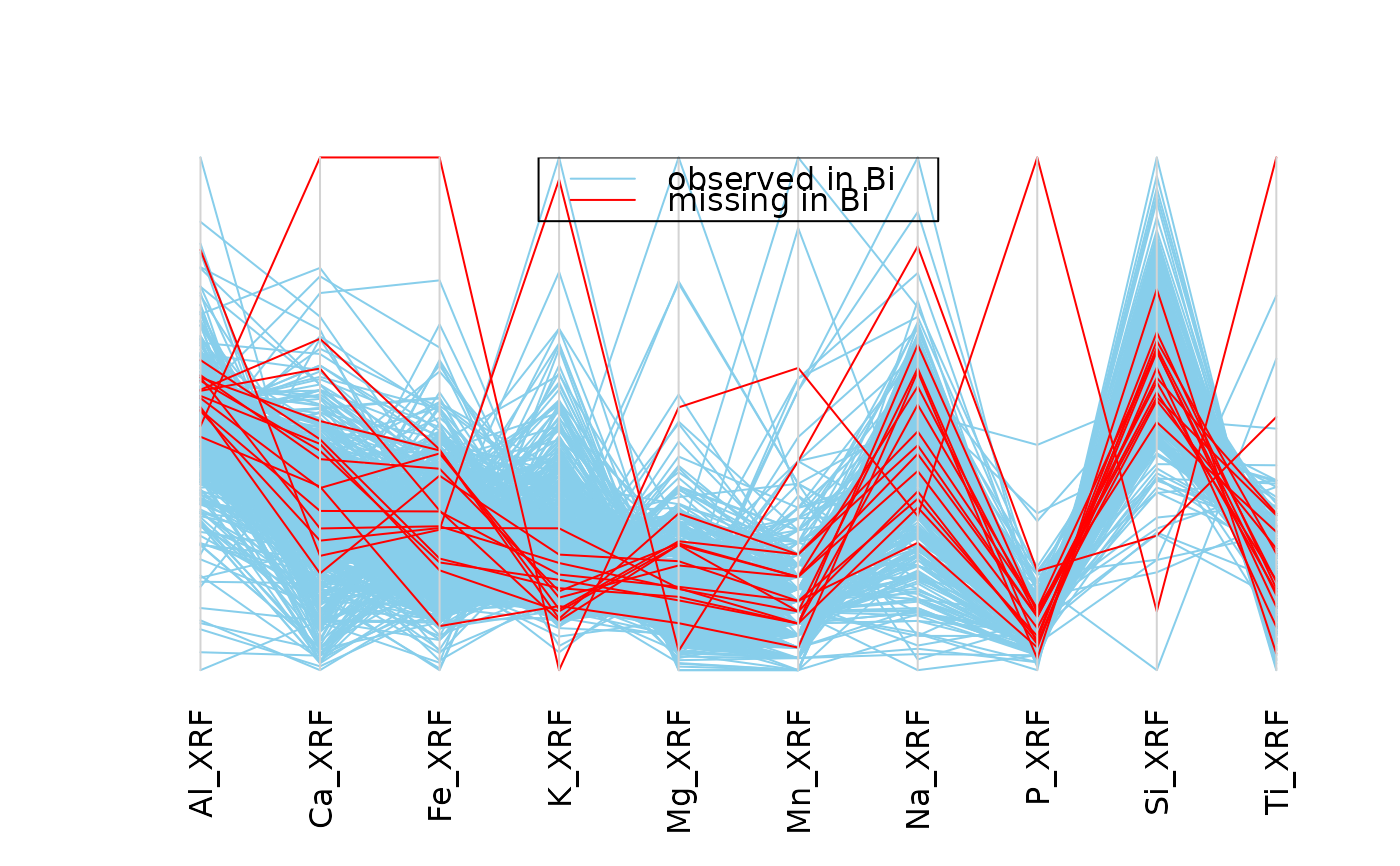
## for missing values
parcoordMiss(chorizonDL[,c(15,101:110)],
plotvars=2:11, interactive = FALSE)
legend("top", col = c("skyblue", "red"), lwd = c(1,1),
legend = c("observed in Bi", "missing in Bi"))
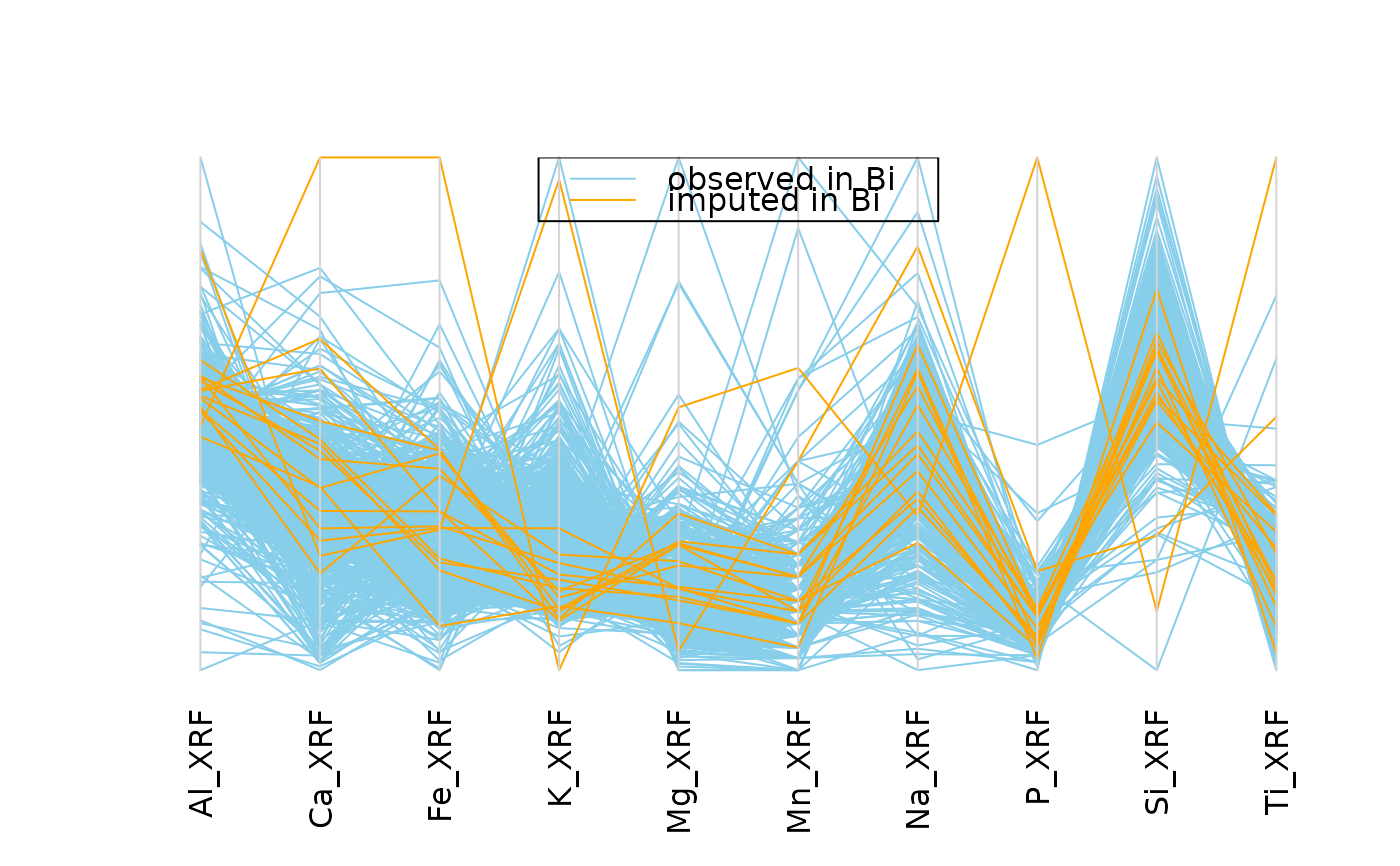
 ## for imputed values
parcoordMiss(kNN(chorizonDL[,c(15,101:110)]), delimiter = "_imp" ,
plotvars=2:11, interactive = FALSE)
legend("top", col = c("skyblue", "orange"), lwd = c(1,1),
legend = c("observed in Bi", "imputed in Bi"))
## for imputed values
parcoordMiss(kNN(chorizonDL[,c(15,101:110)]), delimiter = "_imp" ,
plotvars=2:11, interactive = FALSE)
legend("top", col = c("skyblue", "orange"), lwd = c(1,1),
legend = c("observed in Bi", "imputed in Bi"))