Overview
In addition to Model based Imputation Methods (see
vignette("modelImp")) the VIM package also
presents donor based imputation methods, namely Hot-Deck Imputation,
k-Nearest Neighbour Imputation and fast matching/imputation based on
categorical variable.
This vignette showcases the functions hotdeck() and
kNN(), which can both be used to generate imputations for
several variables in a dataset. Moreover, the function
matchImpute() is presented, which is in contrast a
imputation method based on categorical variables.
Data
The following example demonstrates the functionality of
hodeck() and kNN() using a subset of
sleep. The columns have been selected deliberately to
include some interactions between the missing values.
library(VIM)
dataset <- sleep[, c("Dream", "NonD", "BodyWgt", "Span")]
dataset$BodyWgt <- log(dataset$BodyWgt)
dataset$Span <- log(dataset$Span)
aggr(dataset)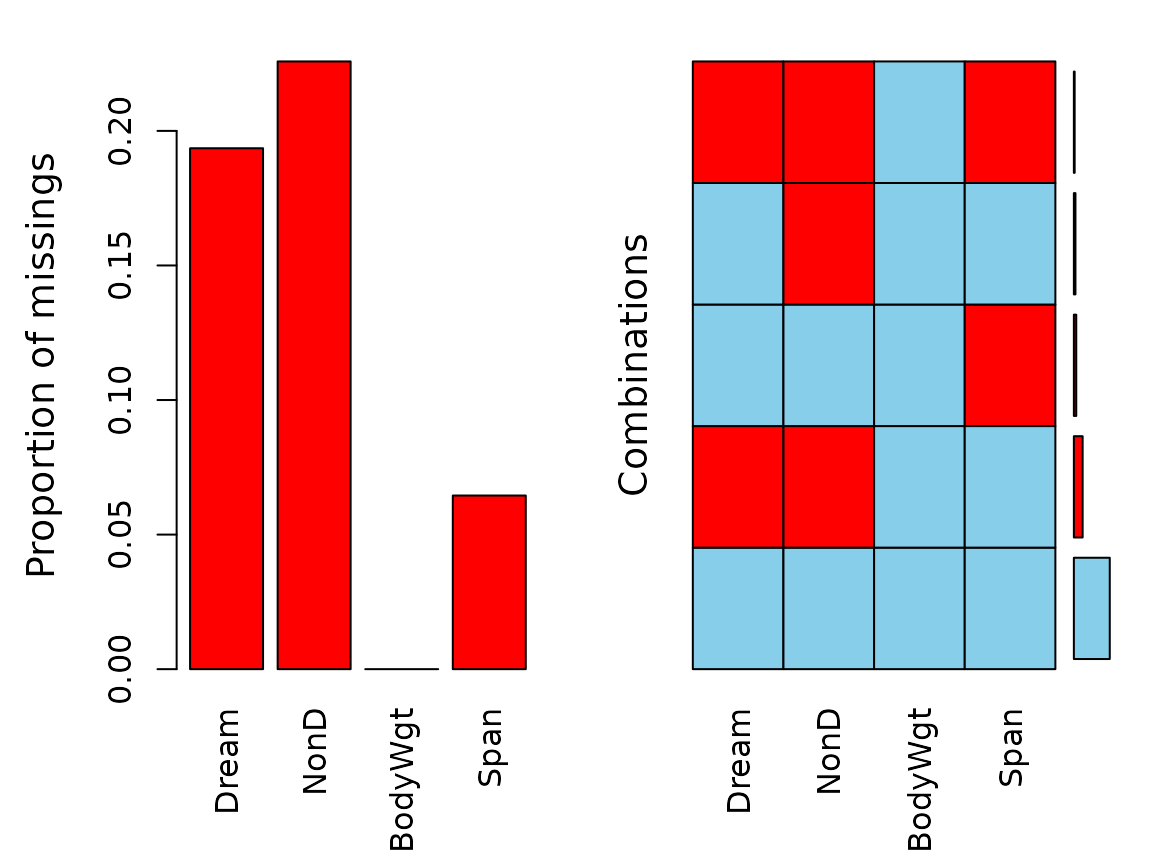
The plot indicates several missing values in Dream,
NonD, and Span.
Imputation
The call of the functions is straightforward. We will start by just
imputing NonD based on the other variables. Besides
imputing missing variables for a single variable, these functions also
support imputation of multiple variables. For matchImpute()
suitable donors are searched based on matching of the categorical
variables.
imp_hotdeck <- hotdeck(dataset, variable = "NonD") # hotdeck imputation
imp_knn <- kNN(dataset, variable = "NonD") # kNN imputation
imp_match <- matchImpute(dataset, variable = "NonD", match_var = c("BodyWgt","Span")) # match imputation
aggr(imp_knn, delimiter = "_imp")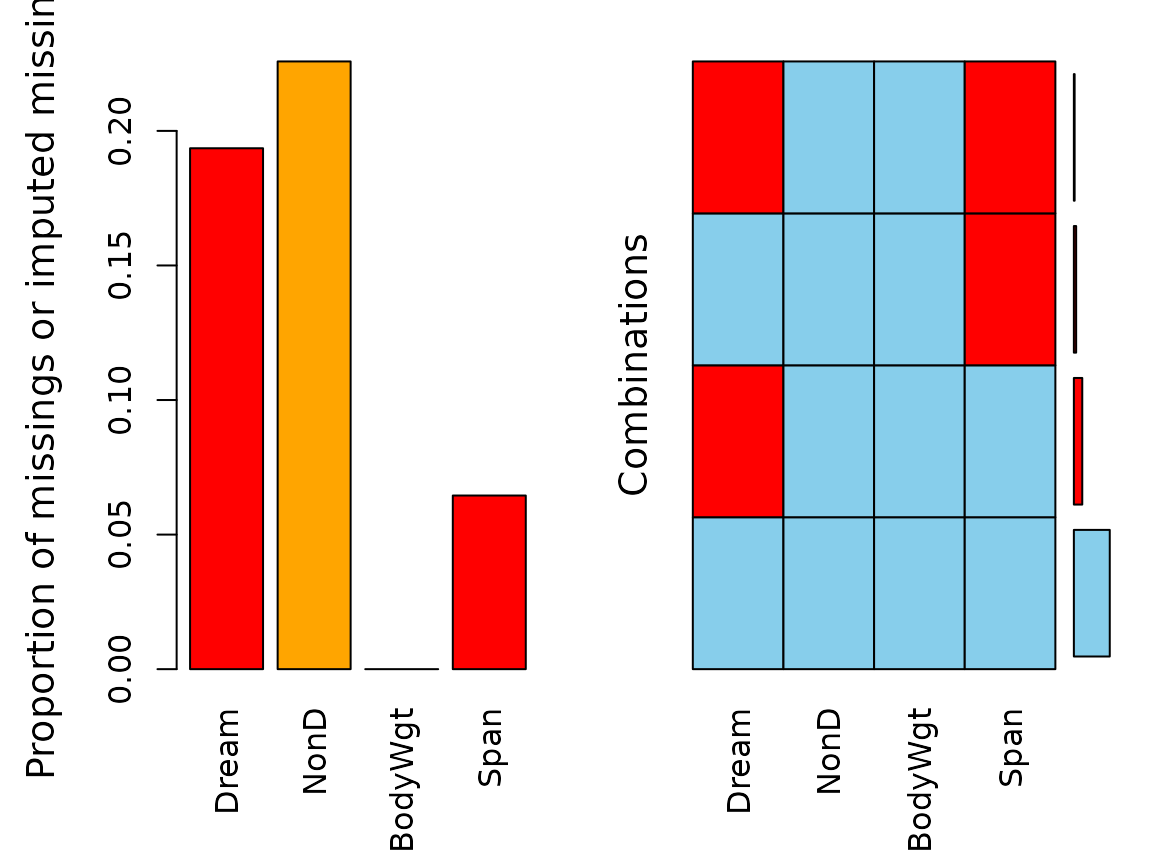
aggr(imp_match, delimiter = "_imp")We can see that kNN() imputed all missing values for
NonD in our dataset. The same is true for the values
imputed via hotdeck(). The specified variables in
matchImpute() serve as a donor and enable imputation for
NonD.
Diagnosing the results
As we can see in the next two plots, the origninal data structure of
NonD and Span is preserved by
hotdeck(). kNN() reveals the typically
procedure of methods, which are based on similar data points weighted by
the distance.
imp_hotdeck[, c("NonD", "Span", "NonD_imp")] |>
marginplot(delimiter = "_imp")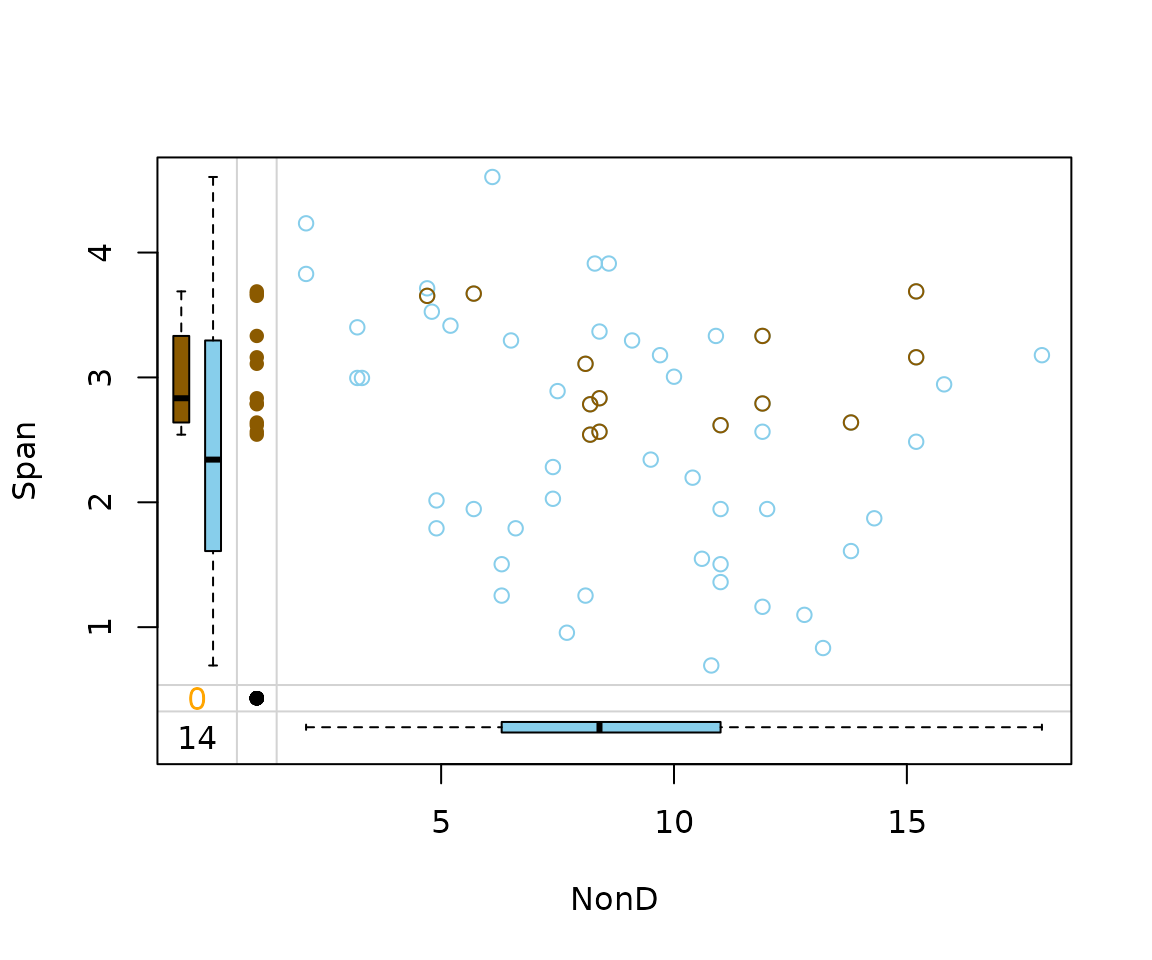
imp_knn[, c("NonD", "Span", "NonD_imp")] |>
marginplot(delimiter = "_imp")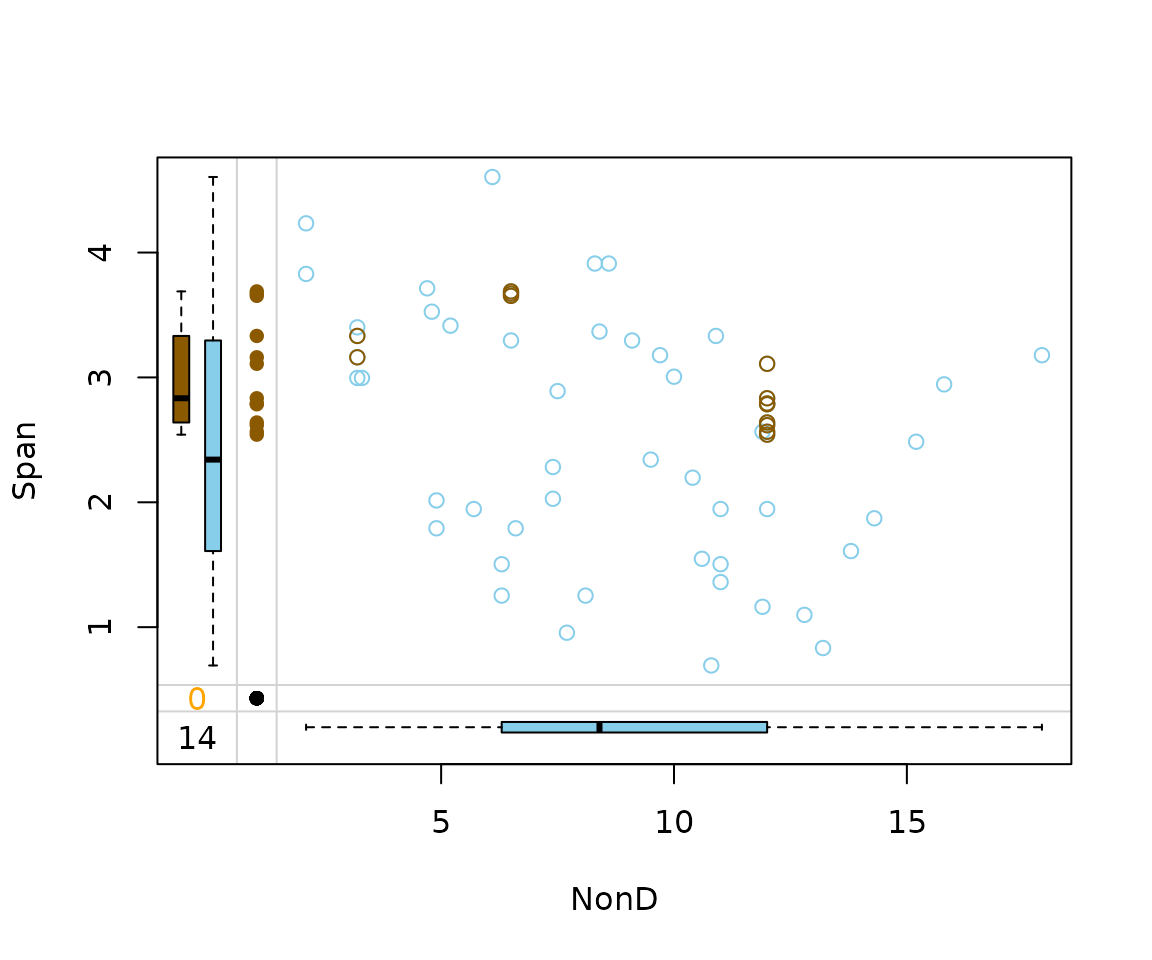
matchImpute() works by sampling values from the suitable
donors and also provides reasonable results.
imp_match[, c("NonD", "Span", "NonD_imp")] |>
marginplot(delimiter = "_imp")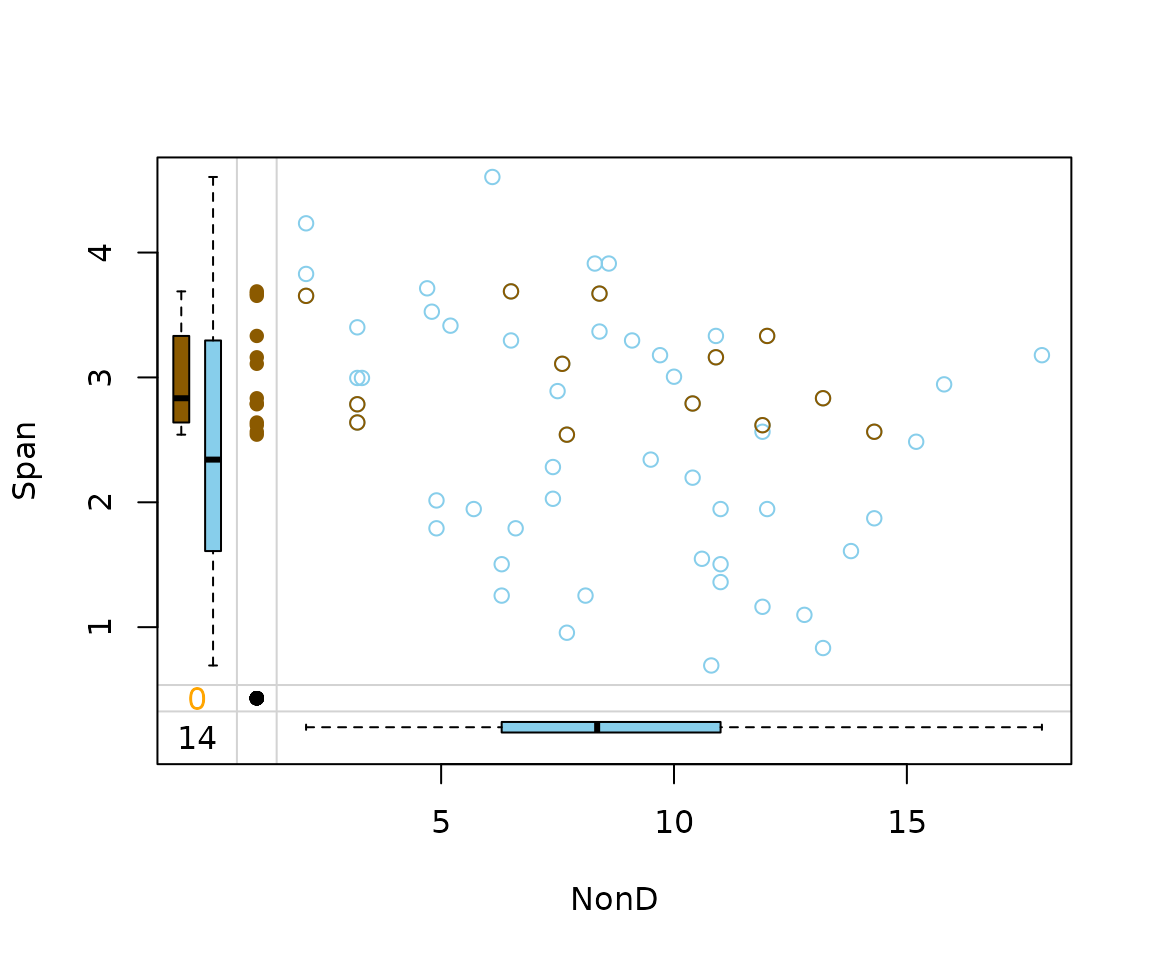
Performance of method
In order to validate the performance of kNN() and to
highlight the ability to impute different datatypes the
iris dataset is used. Firstly, some values are randomly set
to NA.
data(iris)
df <- iris
colnames(df) <- c("S.Length","S.Width","P.Length","P.Width","Species")
# randomly produce some missing values in the data
set.seed(1)
nbr_missing <- 50
y <- data.frame(row = sample(nrow(iris), size = nbr_missing, replace = TRUE),
col = sample(ncol(iris), size = nbr_missing, replace = TRUE))
y<-y[!duplicated(y), ]
df[as.matrix(y)] <- NA
aggr(df)
We can see that there are missings in all variables and some observations reveal missing values on several points.
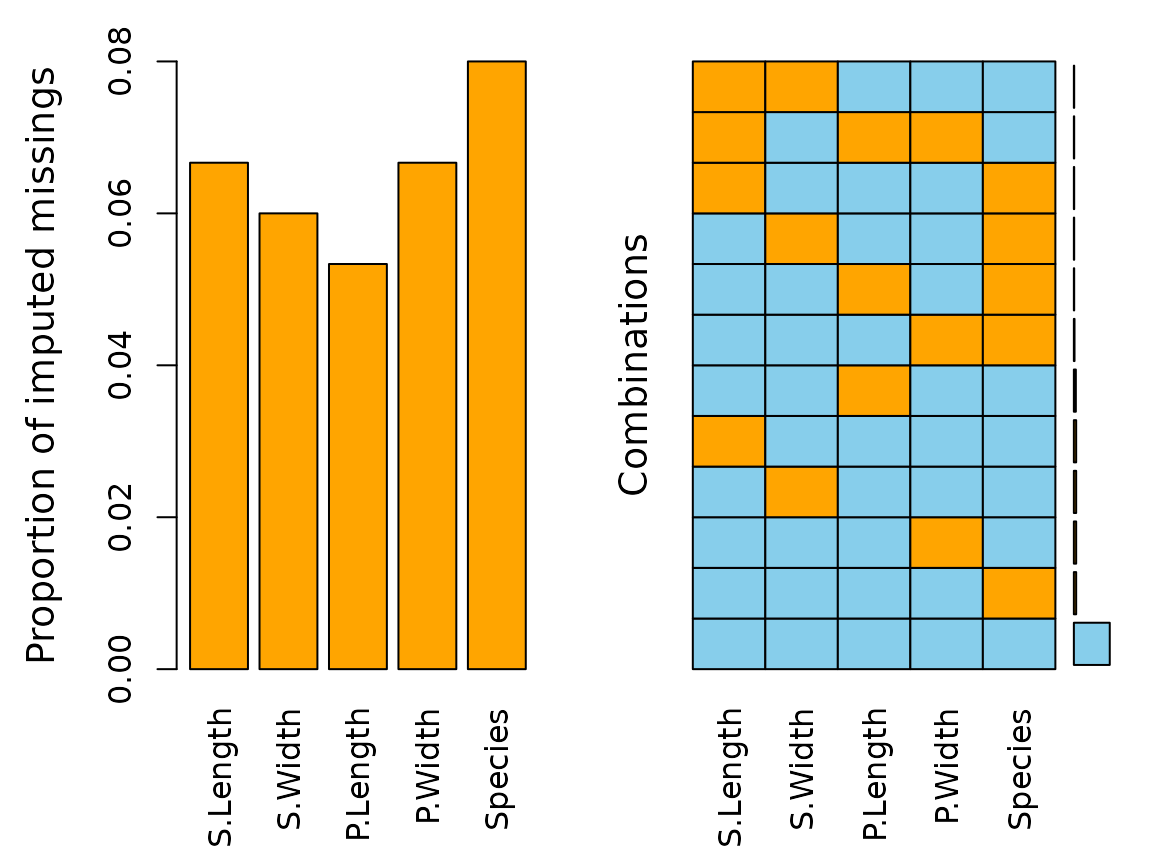
The plot indicates that all missing values have been imputed by
kNN(). The following table displays the rounded first five
results of the imputation for all variables.