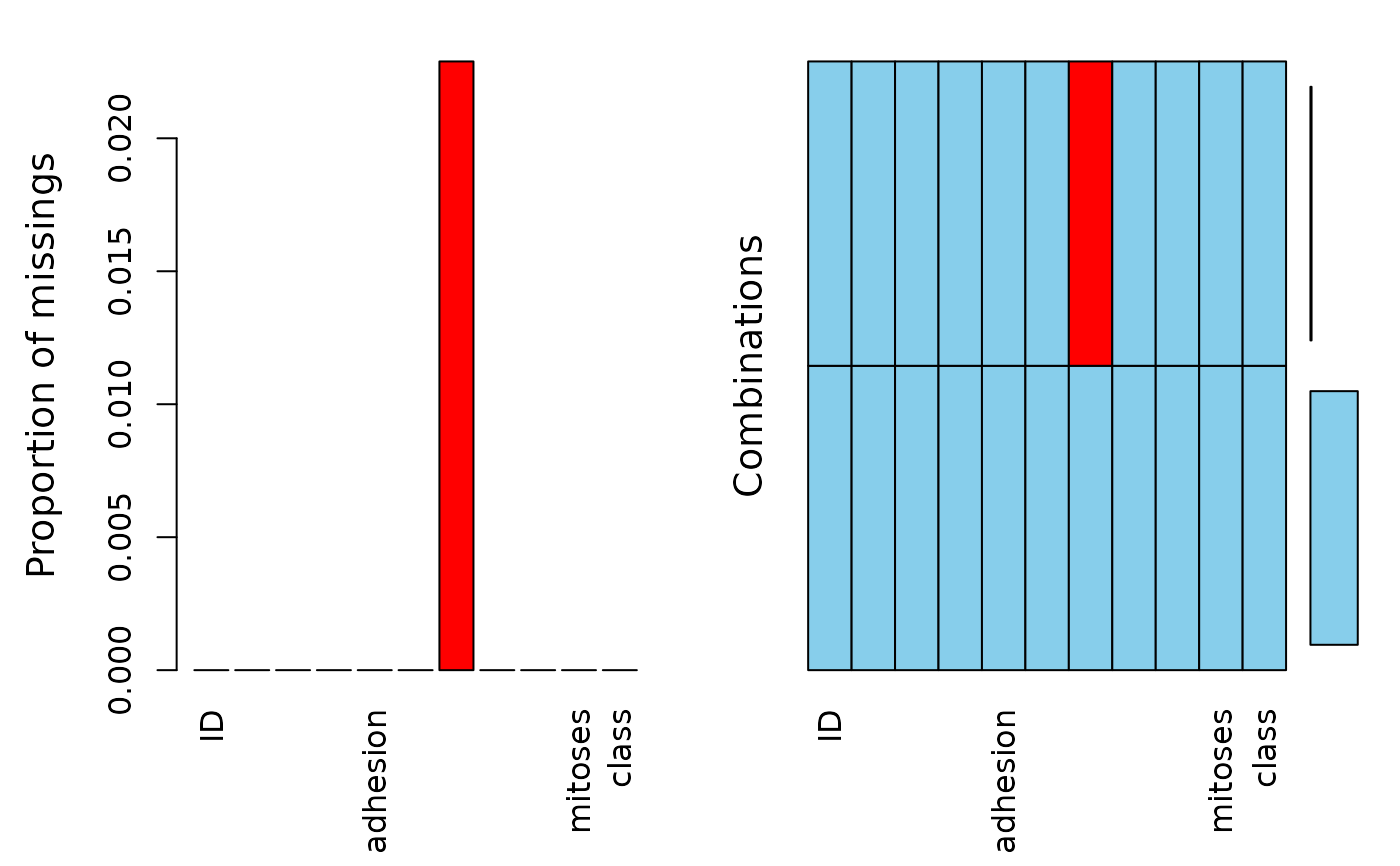
Dataset containing the original Wisconsin breast cancer data.
Format
A data frame with 699 observations on the following 11 variables.
- ID
Sample ID
- clump_thickness
as integer from 1 - 10
- uniformity_cellsize
as integer from 1 - 10
- uniformity_cellshape
as integer from 1 - 10
- adhesion
as integer from 1 - 10
- epithelial_cellsize
as integer from 1 - 10
- bare_nuclei
as integer from 1 - 10, includes 16 missings
- chromatin
as integer from 1 - 10
- normal_nucleoli
as integer from 1 - 10
- mitoses
as integer from 1 - 10
- class
benign or malignant
References
The data downloaded and conditioned for R from the UCI machine learning repository, see https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+(Original) This breast cancer databases was obtained from the University of Wisconsin Hospitals, Madison from Dr. William H. Wolberg. If you publish results when using this database, then please include this information in your acknowledgements. Also, please cite one or more of: O. L. Mangasarian and W. H. Wolberg: "Cancer diagnosis via linear programming", SIAM News, Volume 23, Number 5, September 1990, pp 1 & 18. William H. Wolberg and O.L. Mangasarian: "Multisurface method of pattern separation for medical diagnosis applied to breast cytology", Proceedings of the National Academy of Sciences, U.S.A., Volume 87, December 1990, pp 9193-9196. O. L. Mangasarian, R. Setiono, and W.H. Wolberg: "Pattern recognition via linear programming: Theory and application to medical diagnosis", in: "Large-scale numerical optimization", Thomas F. Coleman and Yuying Li, editors, SIAM Publications, Philadelphia 1990, pp 22-30. K. P. Bennett & O. L. Mangasarian: "Robust linear programming discrimination of two linearly inseparable sets", Optimization Methods and Software 1, 1992, 23-34 (Gordon & Breach Science Publishers).